Protein-coding gene in the species Homo sapiens
| JMJD1C |
|---|
|
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
2YPD, 5FZO |
|
|
| Identifiers |
|---|
| Aliases | JMJD1C, TRIP8, TRIP-8, jumonji domain containing 1C, KDM3C |
|---|
| External IDs | OMIM: 604503; MGI: 1918614; HomoloGene: 3129; GeneCards: JMJD1C; OMA:JMJD1C - orthologs |
|---|
| Gene location (Human) |
|---|
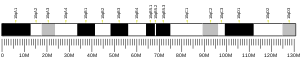
 | | Chr. | Chromosome 10 (human)[1] |
|---|
| | Band | 10q21.3 | Start | 63,167,221 bp[1] |
|---|
| End | 63,521,850 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 10 (mouse)[2] |
|---|
| | Band | 10|10 B5.1 | Start | 67,096,125 bp[2] |
|---|
| End | 67,256,326 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - Achilles tendon
- jejunal mucosa
- superficial temporal artery
- trabecular bone
- right lung
- sperm
- visceral pleura
- tibia
- ganglionic eminence
- monocyte
|
| | Top expressed in | - uterus
- cerebellar cortex
- ganglionic eminence
- neural tube
- ovary
- thymus
- spleen
- mesencephalon
- olfactory bulb
- spermatocyte
|
| | More reference expression data |
|
|---|
| BioGPS | |
|---|
|
| Gene ontology |
|---|
| Molecular function | - oxidoreductase activity
- protein binding
- dioxygenase activity
- metal ion binding
- thyroid hormone receptor binding
- transcription cis-regulatory region binding
- chromatin DNA binding
- histone H3-methyl-lysine-9 demethylase activity
| | Cellular component | - intracellular anatomical structure
- nucleoplasm
- chromatin
- nucleus
| | Biological process | - blood coagulation
- regulation of transcription, DNA-templated
- transcription, DNA-templated
- histone H3-K9 demethylation
- chromatin organization
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | NM_001282948
NM_004241
NM_032776
NM_001318153
NM_001318154
|
|---|
NM_001322252
NM_001322254
NM_001322258 |
| |
|---|
| RefSeq (protein) | NP_001269877
NP_001305082
NP_001305083
NP_001309181
NP_001309183
|
|---|
NP_001309187
NP_116165 |
| |
|---|
| Location (UCSC) | Chr 10: 63.17 – 63.52 Mb | Chr 10: 67.1 – 67.26 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|
Jumonji domain containing 1C is a protein that in humans is encoded by the JMJD1C gene.[5]
Function
The protein encoded by this gene interacts with thyroid hormone receptors and contains a jumonji domain. It is a candidate histone demethylase and is thought to be a coactivator for key transcription factors. It plays a role in the DNA-damage response pathway by demethylating the mediator of DNA damage checkpoint 1 (MDC1) protein, and is required for the survival of acute myeloid leukemia. Mutations in this gene are associated with Rett syndrome and intellectual disability.[5]
Epigenetic regulation of spermatogenesis
Jmjd1C belongs to the Jmjd1 family genes. Jmjd1C encodes a histone H3K9 demethylase. In addition, the JMJD1c gene has a role in mouse spermatogenesis. In male homozygous Jmjd1C mouse knockouts are unable to produce sperm. The mechanism may be the absence of interaction between JMJD1C with JMJD1c's partner proteins, for example, MDC1 and HSP90.[citation needed]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000171988 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000037876 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b "Entrez Gene: Jumonji domain containing 1C".
Further reading
- Grupe A, Li Y, Rowland C, Nowotny P, Hinrichs AL, Smemo S, et al. (2006). "A scan of chromosome 10 identifies a novel locus showing strong association with late-onset Alzheimer disease". Am. J. Hum. Genet. 78 (1): 78–88. doi:10.1086/498851. PMC 1380225. PMID 16385451.
This article incorporates text from the United States National Library of Medicine, which is in the public domain.