| LGMN |
|---|
|
| Available structures |
|---|
| PDB | Ortholog search: PDBe RCSB |
|---|
| List of PDB id codes |
|---|
4N6N, 4N6O, 4FGU, 4AWB, 4AWA, 4AW9 |
|
|
| Identifiers |
|---|
| Aliases | LGMN, AEP, LGMN1, PRSC1, legumain |
|---|
| External IDs | OMIM: 602620; MGI: 1330838; HomoloGene: 38075; GeneCards: LGMN; OMA:LGMN - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 14 (human)[1] |
|---|
| | Band | 14q32.12 | Start | 92,703,807 bp[1] |
|---|
| End | 92,748,679 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 12 (mouse)[2] |
|---|
| | Band | 12|12 E | Start | 102,360,343 bp[2] |
|---|
| End | 102,406,072 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - synovial joint
- gallbladder
- synovial membrane
- right coronary artery
- left lobe of thyroid gland
- placenta
- rectum
- right lobe of thyroid gland
- spleen
- oocyte
|
| | Top expressed in | - yolk sac
- kidney
- calvaria
- saccule
- proximal tubule
- ankle joint
- superior frontal gyrus
- pontine nuclei
- utricle
- primitive streak
|
| | More reference expression data |
|
|---|
| BioGPS | 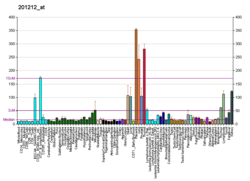 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - cysteine-type peptidase activity
- peptidase activity
- hydrolase activity
- cysteine-type endopeptidase activity
- tau protein binding
| | Cellular component | - late endosome
- apical part of cell
- lysosomal lumen
- lysosome
- extracellular exosome
- endolysosome lumen
- extracellular region
- cytoplasm
- perinuclear region of cytoplasm
| | Biological process | - negative regulation of neuron apoptotic process
- negative regulation of ERBB signaling pathway
- renal system process
- antigen processing and presentation of exogenous peptide antigen via MHC class II
- vitamin D metabolic process
- negative regulation of multicellular organism growth
- vacuolar protein processing
- receptor catabolic process
- proteolysis
- toll-like receptor signaling pathway
- proteolysis involved in cellular protein catabolic process
- response to acidic pH
- memory
- positive regulation of cell population proliferation
- associative learning
- negative regulation of gene expression
- cellular response to hepatocyte growth factor stimulus
- positive regulation of mitotic cell cycle
- cellular response to calcium ion
- positive regulation of monocyte chemotaxis
- dendritic spine organization
- activation of cysteine-type endopeptidase activity
- self proteolysis
- positive regulation of long-term synaptic potentiation
- cellular response to amyloid-beta
- positive regulation of endothelial cell chemotaxis
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | |
|---|
NM_001008530
NM_005606
NM_001363696
NM_001363699 |
| |
|---|
| RefSeq (protein) | |
|---|
NP_001008530
NP_005597
NP_001350625
NP_001350628 |
| |
|---|
| Location (UCSC) | Chr 14: 92.7 – 92.75 Mb | Chr 12: 102.36 – 102.41 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|